QInfer: Bayesian inference for quantum information
Cassandra E. Granade
Centre for Engineered Quantum Systems
$
\newcommand{\ee}{\mathrm{e}}
\newcommand{\ii}{\mathrm{i}}
\newcommand{\dd}{\mathrm{d}}
\newcommand{\id}{𝟙}
\newcommand{\TT}{\mathrm{T}}
\newcommand{\defeq}{\mathrel{:=}}
\newcommand{\Tr}{\operatorname{Tr}}
\newcommand{\Var}{\operatorname{Var}}
\newcommand{\Cov}{\operatorname{Cov}}
\newcommand{\rank}{\operatorname{rank}}
\newcommand{\expect}{\mathbb{E}}
\newcommand{\sket}[1]{|#1\rangle\negthinspace\rangle}
\newcommand{\sbraket}[1]{\langle\negthinspace\langle#1\rangle\negthinspace\rangle}
\newcommand{\Gini}{\operatorname{Ginibre}}
\newcommand{\supp}{\operatorname{supp}}
\newcommand{\ket}[1]{\left|#1\right\rangle}
\newcommand{\bra}[1]{\left\langle#1\right|}
\newcommand{\braket}[1]{\left\langle#1\right\rangle}
$
joint work with Christopher Ferrie
contributions from
Steven Casagrande, Ian Hincks, Jonathan Gross, Michal Kononenko, Thomas Alexander, and Yuval Sanders
Characterization plays a number of different roles in
quantum information experiments.
- Calibration
- (Rabi/Ramsey/phase est., crosstalk learning)
- Diagnosis and Debugging
- (Tomography)
- Verification and Validation
- (RB, quantum Hamiltonian learning)
All of these are examples of parameter estimation.
## Parameter Estimation ##
Given data $D$, and a model $\vec{x}$,
what should we estimate $\vec{x}$ as?
- Rabi/Ramsey: $\vec{x} = (\omega)$
- Crosstalk/Hamiltonian learning: $\vec{x} = \operatorname{vec}(H)$
- Tomography: $\vec{x} = \operatorname{vec}(\rho)$
---
From an experimental perspective, parameter estimation
isn't the point, but a tool to get things done.
### **Example**: Ramsey Estimation ###
Suppose $H = \omega \sigma_z / 2$ for some unknown $\omega$.
To learn $\omega$:
- Prepare $\ket{+} \propto \ket{0} + \ket{1}$, measure “click” w/ pr.:
$
\|\bra{+} \ee^{\ii \omega t \sigma_z / 2} \ket{+}\|^2 = \cos^2(\omega t / 2)
$.
- Repeat for many “shots” to estimate click pr.
- Repeat for many times to estimate signal.
You'll get something that looks a bit like this:
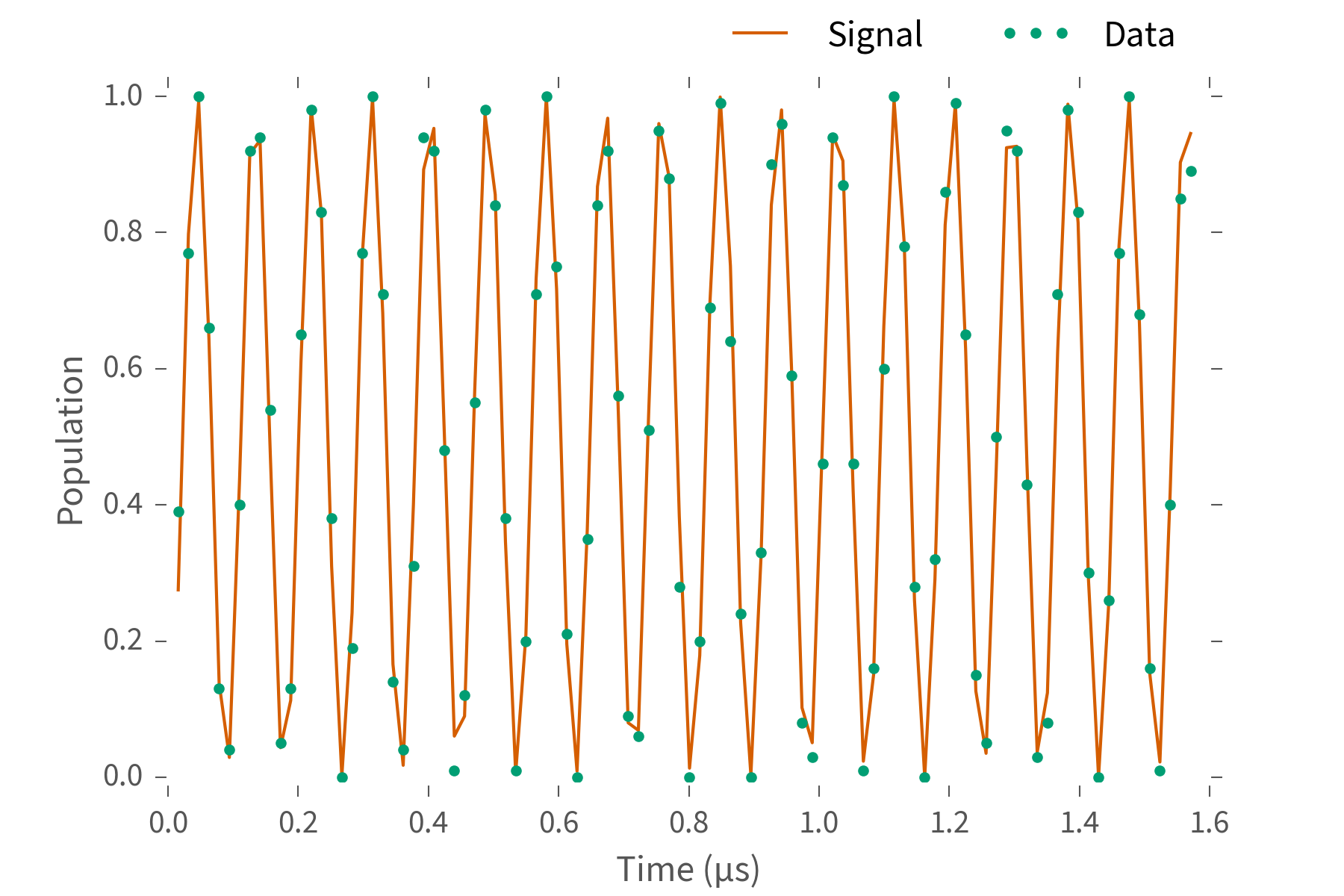 What's $\omega$? Fourier transform and look at the peak.
What's $\omega$? Fourier transform and look at the peak.
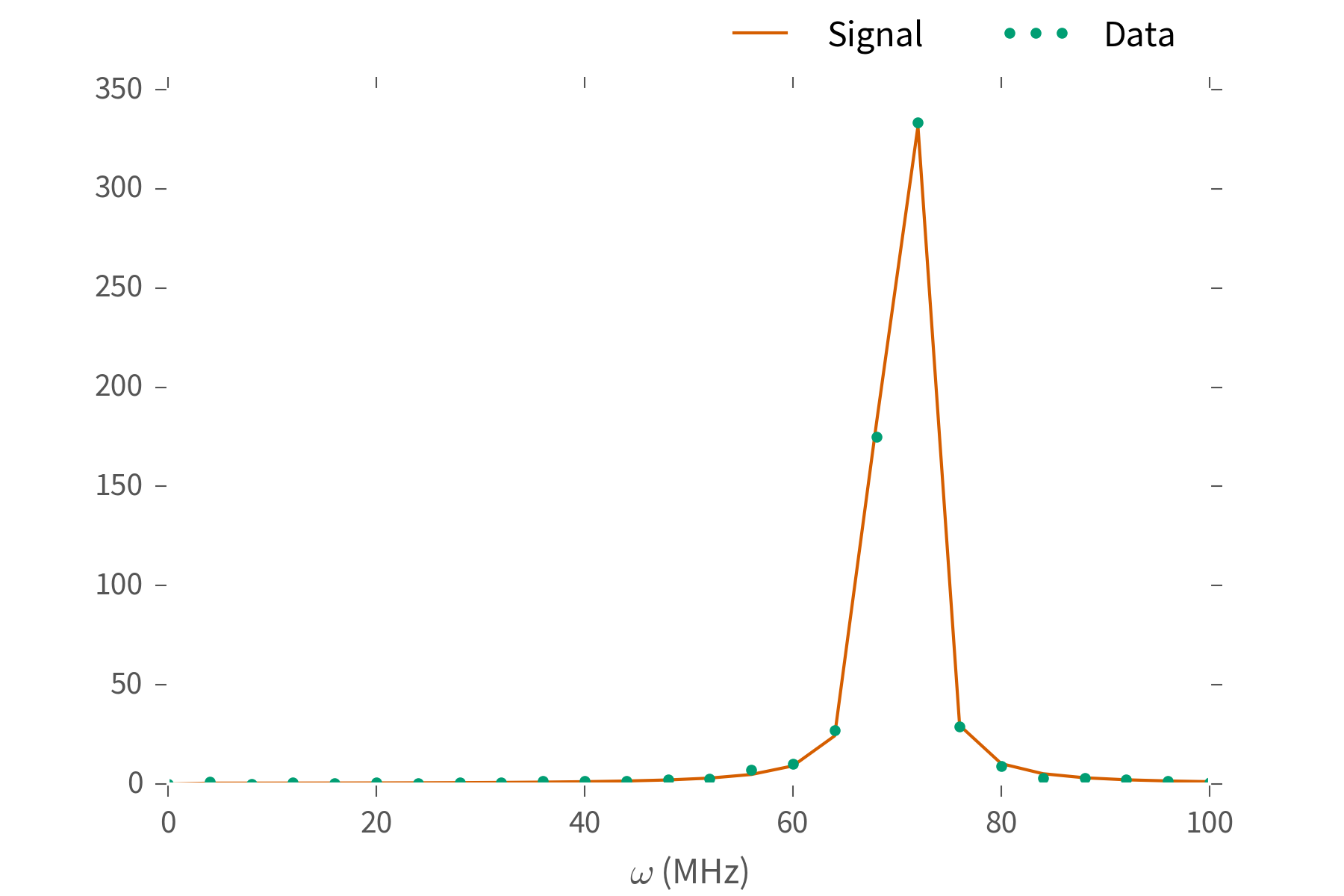 We can do better.
We can do better.
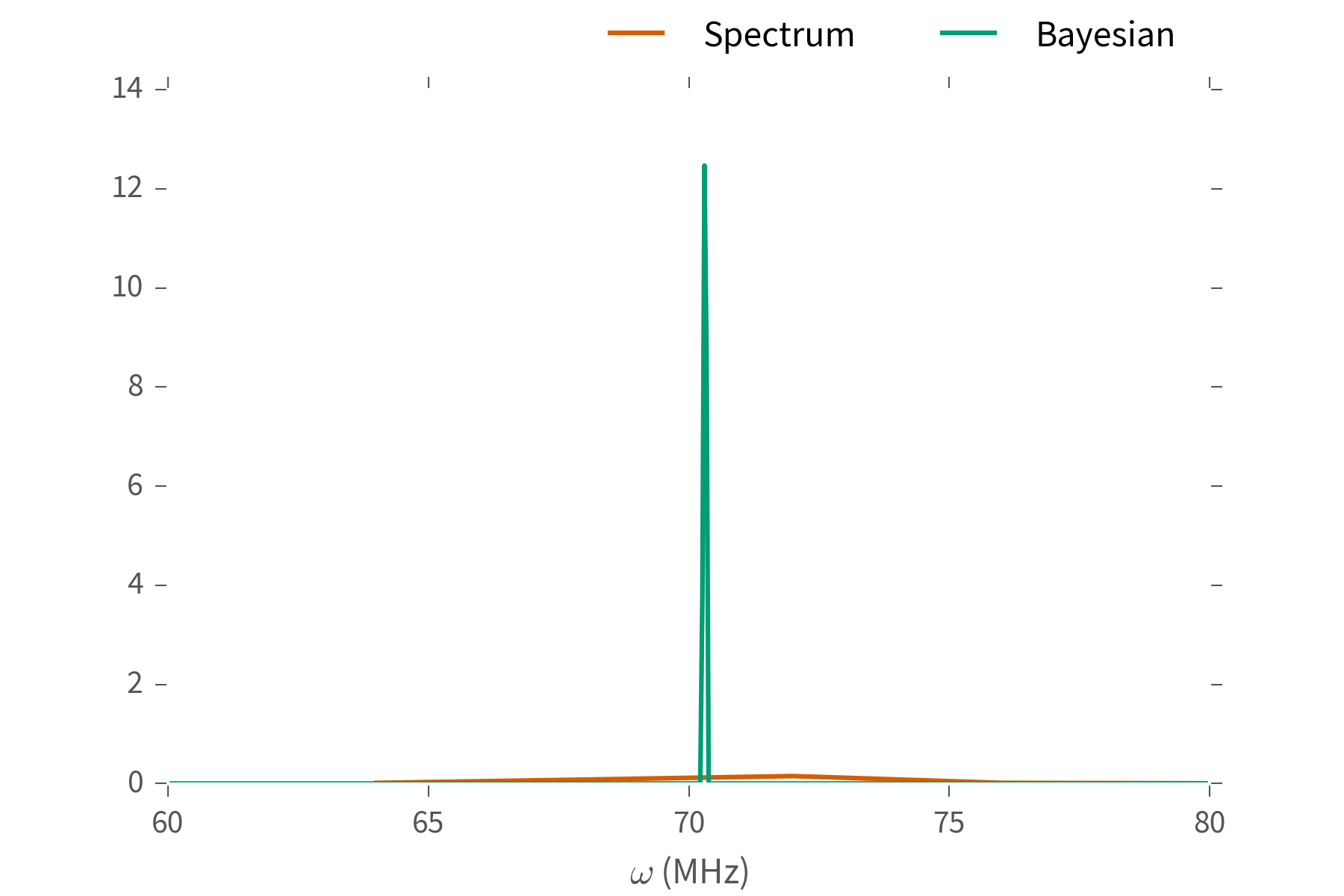
Our goal is to make useful tools for parameter estimation
that work in practice, in a statistically-principled
fashion.
Make it easier to get experiments done:
- Reduce data collection costs
- Provide accurate estimates
- Support reproducible research practices.
...but first, an aside on scientific software in quantum information.
Our field is very well publicly supported.
Thus, our science should support the public.
- **Reproducible**: open data, source
- **Accessible**: e.g. green/gold
Perhaps most of all, we must make sure that our
results are **reusable**.
---
Our work should enable further research,
not prevent it.
This flies in the face of how physicists are taught to program
and present numerical results.
*(e.g. dependent on closed software, no data, source or documentation provided)*
Let's do **reproducible**, **accessible** and **reusable**
research instead.
Our theoretical basis will be
Bayesian Inference with Particle Filtering.
Represent our beliefs about the model by a set of
hypotheses $\{\vec{x}_i\}$, along with their weights
$\{w_i\}$.
\begin{align*}
\text{Estimate: } \hat{x} & = \sum_i w_i \vec{x}_i \\
\text{Update: } w_i' & \propto w_i \times \Pr(D | \vec{x}_i)
\end{align*}
Numerical stability is provided by resampling:
- Contract hypotheses towards center of mass
- Convolve with Gaussian
Preserves estimates and errors of hypotheses $\{\vec{x}\}$,
while restoring stability of the approximation.
- Statistically principled: approximates Bayesian
posteriors, makes dependence on prior *explicit*.
- Very general *and extensible* approach.
- Provides rich error reporting, model selection and other
diagnostics.
>>> import qinfer
Implements particle filtering, with support for common
quantum information models:
- Hamiltonian learning / phase estimation
- Randomized benchmarking
- State and process tomography
### Scientific Software is for People, not Computers ###
Thus, **QInfer** is:
- **Accessible**: written in Python,
usable from Python, Julia, MATLAB.
- **Open source**: modifiable and reproducible.
- **Portable**: Windows/Linux/OS X.
- **Legible**: well-documented (guide & examples).
- **Expressive**: represents the scientific concepts.
### Installing **QInfer** ###
```bash
$ pip install qinfer
```
Works on Python 2.7, 3.3, 3.4, and 3.5 with the Anaconda
Distribution.
---
For Julia, need one additional step:
```julia
julia> Pkg.add("PyCall")
```
### Using **QInfer**: Simple Estimation ###
Make the data...
```python
import numpy as np
true_omega = 70.3
n_shots = 100
ts = np.pi * np.arange(1, 101) / (2 * 100.0);
signal = np.sin(true_omega * ts / 2) ** 2;
counts = np.random.binomial(n=n_shots, p=ideal_signal)
```
...then process it.
```python
import qinfer as qi
data = np.column_stack([counts, ts, n_shots * np.ones(len(ts))])
est_mean, est_cov = qi.simple_est_prec(data, freq_min=0, freq_max=100)
```
### Using **QInfer** from MATLAB 2016a ###
Same idea: make the data...
```matlab
true_omega = 70.3;
n_shots = 400;
ts = pi * (1:1:100) / (2 * 100);
signal = sin(true_omega * ts / 2) .^ 2;
counts = binornd(n_shots, signal);
```
... then process it.
```matlab
setenv MKL_NUM_THREADS 1
data = py.numpy.column_stack({counts ts ...
n_shots * ones(1, size(ts, 2))});
est = py.qinfer.simple_est_prec(data, ...
pyargs('freq_min', 0, 'freq_max', 100));
```
### Using **QInfer** from Julia ###
Same for Julia: make the data...
```julia
true_omega = 70.3
n_shots = 100
ts = pi * (1:1:100) / (2 * 100)
signal = sin(true_omega * ts / 2) .^ 2
counts = map(p -> rand(Binomial(n_shots, p)), signal);
```
...then process it.
```julia
@pyimport numpy as np
@pyimport qinfer as qi
data = [counts'; ts'; n_shots * ones(length(ts))']'
est_mean, est_cov = qi.simple_est_prec(data, freq_min=0, freq_max=100)
```
### Breaking it Down ###
**QInfer** is built up of several main components:
- ``Model``: Specifies a model for what parameters describe
an experiment.
- ``Distribution``: Specifies what is known about those
parameters at the start.
- ``SMCUpdater``: Uses sequential Monte Carlo to update
knowledge based on data.
- ``Heuristic``: Selects new experiments to perform.
### **QInfer** Concepts: Models ###
Parameter esimation problems are specified as **models**,
defining parameters of interest, what data looks like, etc.
```python
>>> SimplePrecessionModel()
>>> BinomialModel(RandomizedBenchmarkingModel())
>>> BinomialModel(TomographyModel(basis))
```
### **QInfer** Concepts: Models ###
Models expose two very important functionalities:
- ``simulate_experiment``: Simulates data $d$
from an experiment $e$, according
to a set of model parameters $\vec{x}$.
- ``likelihood``: Returns the probability $\Pr(d | \vec{x}; e)$
of observing $d$ in an experiment $e$
given model parameters $\vec{x}$.
The ``likelihood`` function returns a *tensor*
$$
L_{ijk} = \Pr(d_i | \vec{x}_j; e_k).
$$
```python
L = SimplePrecessionModel().likelihood(
1, # outcomes
np.array([w]), # models
ts # experiments
)
```
Enables:
- Vectorization over data, models and experiments
- Parallelization
- Caching of intermediate results
# Index along data and models to get a vector over experiments.
plt.plot(ts, L[0, 0, :])
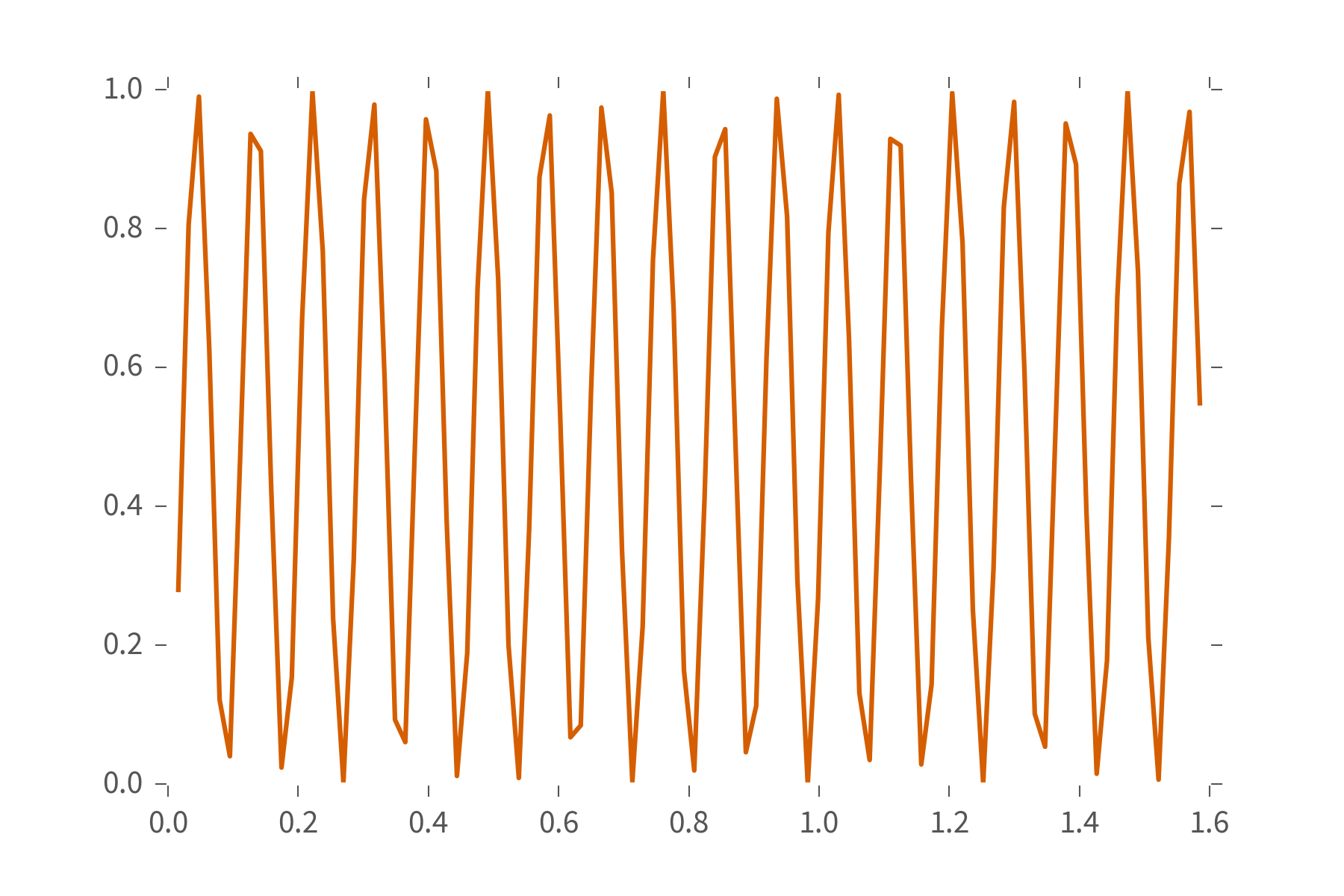 ### **QInfer** Concepts: Distributions ###
A `Distribution` is an object that produces *samples*.
**QInfer** comes with several built-in...
```python
>>> UniformDistribution([0, omega_max])
>>> plt.hist(NormalDistribution(0, 2).sample(n=100000), bins=20)
```
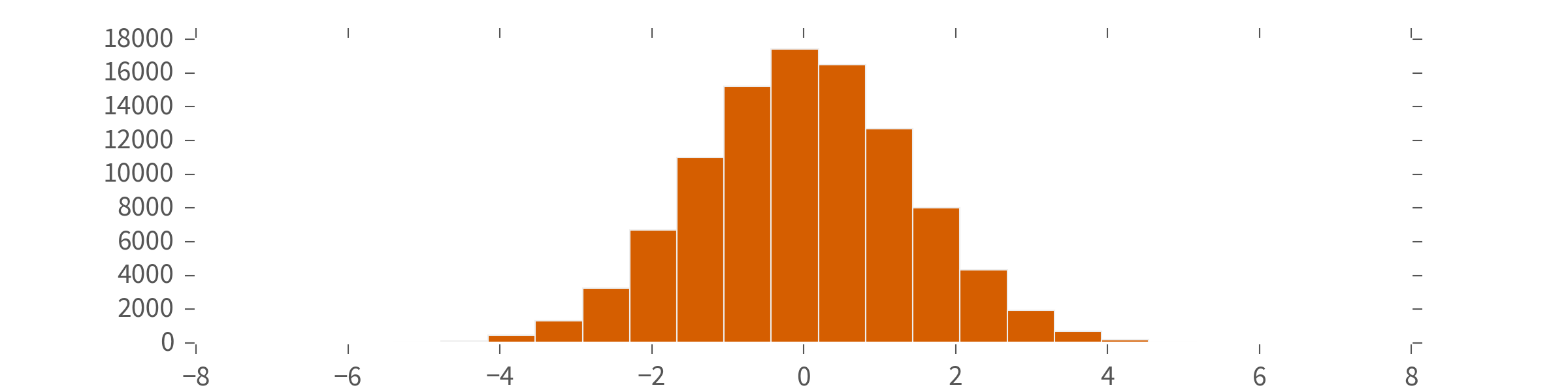
### **QInfer** Concepts: Distributions ###
A `Distribution` is an object that produces *samples*.
**QInfer** comes with several built-in...
```python
>>> UniformDistribution([0, omega_max])
>>> plt.hist(NormalDistribution(0, 2).sample(n=100000), bins=20)
```

### **QInfer** Concepts: Distributions ###
...including exotic distributions such as redit priors.
```python
>>> plot_rebit_prior(
... GinibreReditDistribution(pauli_basis(1)),
... rebit_axes=[1, 3]
... )
```
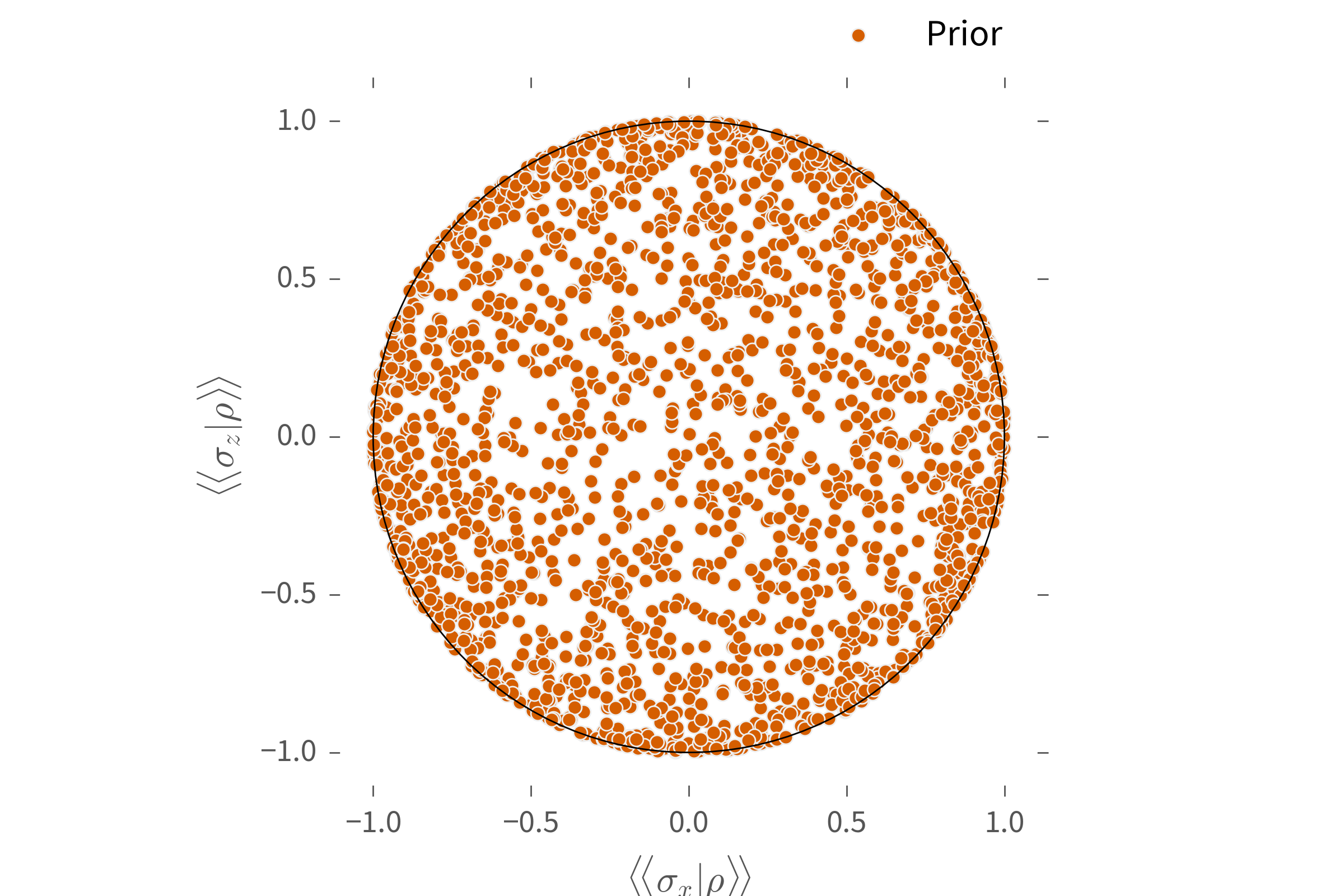 ### **QInfer** Concepts: Distributions ###
Distributions can also be combined in different ways:
```python
>>> ProductDistribution(
... NormalDistribution([0.9, 0.1 ** 2]),
... UniformDistribution([0, 1]),
... ConstantDistribution(0)
... )
```
### Using **QInfer**: Updating ###
Typically, once you have a model and a prior, learning
parameters then proceeds by looping over data:
```python
>>> updater = SMCUpdater(model, n_particles, prior)
>>> for idx in range(n_measurements):
... experiment = # select the next experiment
... datum = # make a measurement
... updater.update(datum, experiment)
>>> est = updater.est_mean()
```
### Using **QInfer**: Updating ###
The updated distribution provides estimates,
error bars, and plots.
#### **Example**: Randomized Benchmarking ####
```python
>>> mean, cov, extra = qi.simple_est_rb(
... # Use return_all=True to get the updater.
... data, p_min=0.8, return_all=True
... )
>>> print(mean[0], "±", np.sqrt(cov)[0, 0])
0.991188359708 ± 0.0012933975599
>>> extra['updater'].plot_posterior_marginal(idx_param=0)
>>> extra['updater'].plot_covariance(corr=True)
```
### **QInfer** Concepts: Heuristics ###
*Heuristics* can be used to design measurements.
For example, $t_k = ab^k$ is optimal for non-adaptive
Rabi/Ramsey/phase estimation.
```
>>> heuristic = ExpSparseHeuristic(scale=a, base=b)
```
**QInfer** implements heuristics as functions which provide
new experiments.
### Updater Loops Revisited ###
For instance, using a heuristic ``heuristic_class`` and
simulating data, we can flesh out the updater loop.
```python
>>> updater = SMCUpdater(model, n_particles, prior)
>>> heuristic = heuristic_class(updater)
>>> for idx in range(n_measurements):
... experiment = heuristic()
... datum = model.simulate_experiment(true_model, experiment)
... updater.update(datum, experiment)
>>> est = updater.est_mean()
```
### Tomography Updater Loop ###
```python
from qinfer.tomography import *
basis = pauli_basis(1)
prior = GinibreReditDistribution(basis)
model = BinomialModel(TomographyModel(basis))
updater = SMCUpdater(model, 2000, prior)
heuristic = RandomPauliHeuristic(updater,
other_fields={'n_meas': 40}
)
for idx_exp in xrange(50):
experiment = heuristic()
datum = model.simulate_experiment(true_state, experiment)
updater.update(datum, experiment)
```
### **QInfer** Concepts: Distributions ###
Distributions can also be combined in different ways:
```python
>>> ProductDistribution(
... NormalDistribution([0.9, 0.1 ** 2]),
... UniformDistribution([0, 1]),
... ConstantDistribution(0)
... )
```
### Using **QInfer**: Updating ###
Typically, once you have a model and a prior, learning
parameters then proceeds by looping over data:
```python
>>> updater = SMCUpdater(model, n_particles, prior)
>>> for idx in range(n_measurements):
... experiment = # select the next experiment
... datum = # make a measurement
... updater.update(datum, experiment)
>>> est = updater.est_mean()
```
### Using **QInfer**: Updating ###
The updated distribution provides estimates,
error bars, and plots.
#### **Example**: Randomized Benchmarking ####
```python
>>> mean, cov, extra = qi.simple_est_rb(
... # Use return_all=True to get the updater.
... data, p_min=0.8, return_all=True
... )
>>> print(mean[0], "±", np.sqrt(cov)[0, 0])
0.991188359708 ± 0.0012933975599
>>> extra['updater'].plot_posterior_marginal(idx_param=0)
>>> extra['updater'].plot_covariance(corr=True)
```
### **QInfer** Concepts: Heuristics ###
*Heuristics* can be used to design measurements.
For example, $t_k = ab^k$ is optimal for non-adaptive
Rabi/Ramsey/phase estimation.
```
>>> heuristic = ExpSparseHeuristic(scale=a, base=b)
```
**QInfer** implements heuristics as functions which provide
new experiments.
### Updater Loops Revisited ###
For instance, using a heuristic ``heuristic_class`` and
simulating data, we can flesh out the updater loop.
```python
>>> updater = SMCUpdater(model, n_particles, prior)
>>> heuristic = heuristic_class(updater)
>>> for idx in range(n_measurements):
... experiment = heuristic()
... datum = model.simulate_experiment(true_model, experiment)
... updater.update(datum, experiment)
>>> est = updater.est_mean()
```
### Tomography Updater Loop ###
```python
from qinfer.tomography import *
basis = pauli_basis(1)
prior = GinibreReditDistribution(basis)
model = BinomialModel(TomographyModel(basis))
updater = SMCUpdater(model, 2000, prior)
heuristic = RandomPauliHeuristic(updater,
other_fields={'n_meas': 40}
)
for idx_exp in xrange(50):
experiment = heuristic()
datum = model.simulate_experiment(true_state, experiment)
updater.update(datum, experiment)
```
## **QInfer** 💖 **QuTiP** ##
Tomography support in **QInfer** is backed by
the quantum object representation and prior
distributions
provided by **QuTiP**
3.2.
```python
>>> modelparams = basis.state_to_modelparams(rho)
>>> rho = basis.modelparams_to_state(rho)
>>> cov_superop = basis.covariance_mtx_to_superop(
... updater.est_covariance_mtx()
... )
```
Makes it easy to integrate with other QIP concepts: fidelity, Schatten and diamond norms, etc.
### **QInfer** Concepts: Performance Testing ###
In both of these examples, we assumed that the true model
was known. This lets us quickly assess how well **QInfer**
works for a given model.
```python
>>> performance = perf_test_multiple(
... n_trials=400,
... model=BinomialModel(SimplePrecessionModel()),
... n_particles=2000,
... prior=UniformDistribution([0, 1]),
... n_exp=200,
... # partial lets us quickly make new heuristic classes.
... heuristic_class=partial(
... ExpSparseHeuristic, other_fields={'n_meas': 40}
... )
... )
```
### Simple Parallelization ###
First, we connect to our engines...
```python
from ipyparallel import *
c = Client()
load_balanced_view = c.load_balanced_view()
```
...then we can easily delegate trials.
```python
basis = pauli_basis(1)
performance = perf_test_multiple(
n_trials=400,
model=BinomialModel(TomographyModel(basis)),
n_particles=2000,
prior=GinibreDistribution(basis),
n_exp=100,
heuristic_class=partial(
RandomPauliHeuristic, other_fields={'n_meas': 40}
),
apply=load_balanced_view.apply, # ← parallel!
progressbar=IPythonProgressBar # ← Jupyter!
)
```
### Diffusive Models ###
**QInfer** also supports time-dependent parameter estimation
by adding an update rule to hypothesis positions as well as
weights:
$$
\vec{x}(t_{k + 1}) - \vec{x}(t_k) \sim \mathcal{N}(0, \sigma^2).
$$
Diffusive estimation can still work even if the underlying trajectory
is deterministic.
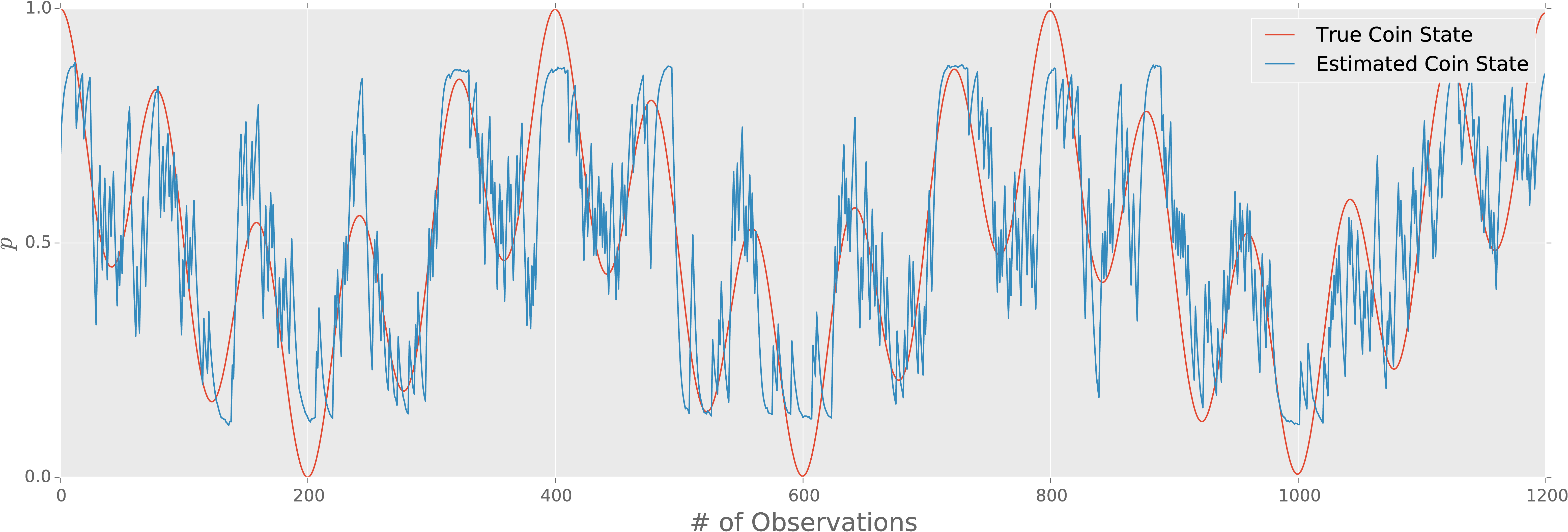
For example, suppose a coin bias evolves as $\Pr(\text{heads}) = p = \frac12 (\cos^2(\omega_1 t / 2) + \cos^2(\omega_2 t) / 2))$.
How can we use QInfer to do reproducible
research?
Practical adaptive quantum tomography
1605.05039
• joint work with Chris Ferrie and Steve Flammia
Self-guided tomography (Ferrie 10/bchr) as a heuristic
for adaptive tomography.
- Based on QInfer and QuTiP.
- All figures generated from a single Jupyter Notebook.
- Source and data available on Figshare 10/bhfk.
Our hope is that **QInfer** will thus be a useful tool for theory and experiment alike.
---
- Source: https://github.com/QInfer/python-qinfer
- Docs: http://docs.qinfer.org
- Try it out online: https://goo.gl/zWt9Du
*Version 1.0 coming soon.*






